The Biological Fortune Teller: Why Morphology is the New Frontier of Biologics
In the high-stakes world of drug development, the most valuable commodity is not just the genetic sequence of a new antibody, but the time it takes to produce it. For decades, the process of Cell Line Development (CLD) has been a study in patience: scientists deposit single cells into thousands of wells and wait weeks to see which ones grow, which ones produce, and which ones are “monoclonal” enough to satisfy regulators.
But a series of recent breakthroughs suggests that the industry is entering a new era. We are moving away from waiting for biological growth and toward predicting it through the lens of high-resolution imaging and artificial intelligence.
The 36-Day Sprint
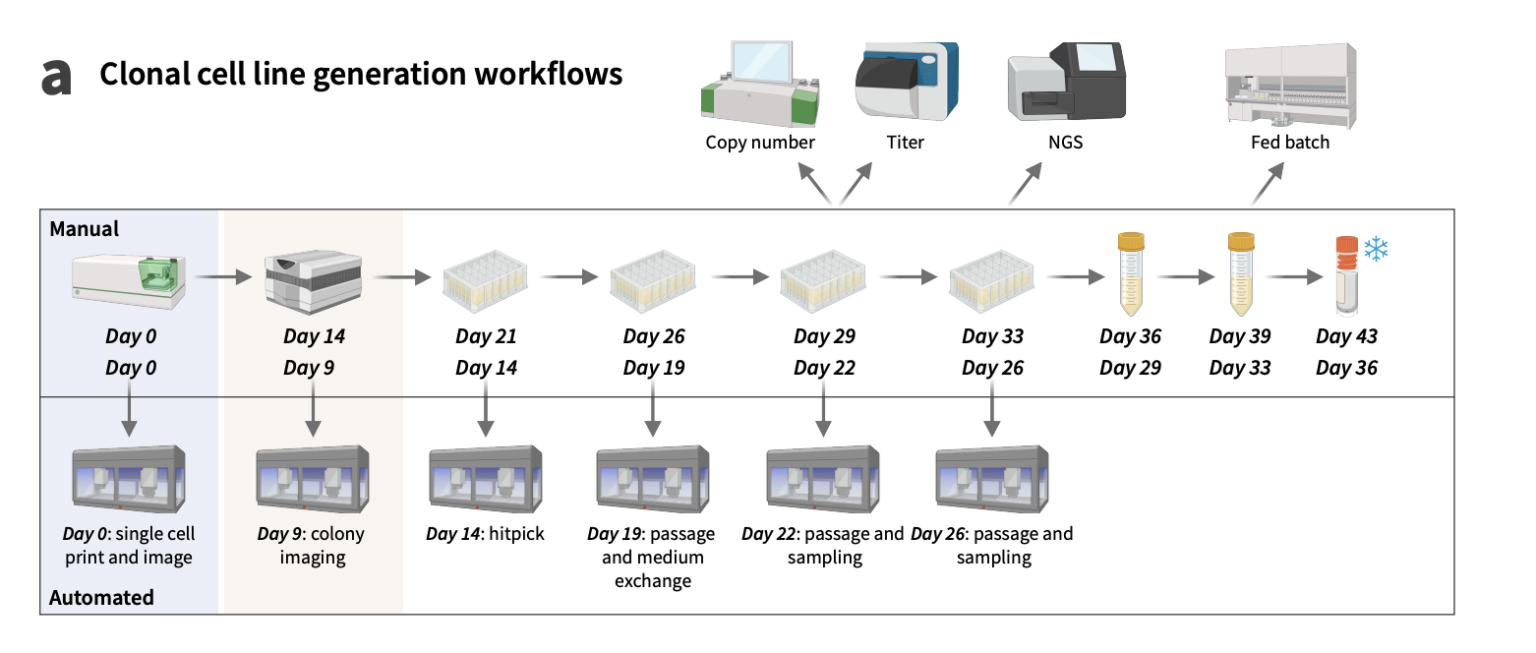
The baseline for efficiency was recently reset by researchers at Gilead Sciences. Traditionally, a manual CLD workflow required roughly 43 days of labor-intensive culture and imaging. By introducing CLAIRE [2] (Cell Line AI Recognition and Evaluation), a tool that integrates deep-learning image analysis with robotic liquid handling, the team compressed this timeline to just 36 days.
CLAIRE [2]: By integrating the CLAIRE toolkit with automated liquid handling, researchers compressed the standard 43-day manual cell line development (CLD) cycle into a brisk 36-day sprint.
At the heart of CLAIRE is the DETR (Detection Transformer) model, which achieved an F1-score of over 0.90 in identifying single cells, the “holy grail” of regulatory compliance. By moving monoclonality verification from a subjective manual task to an automated digital output, the workflow significantly increased daily throughput while reducing the risk of human error.
Fate at First Sight
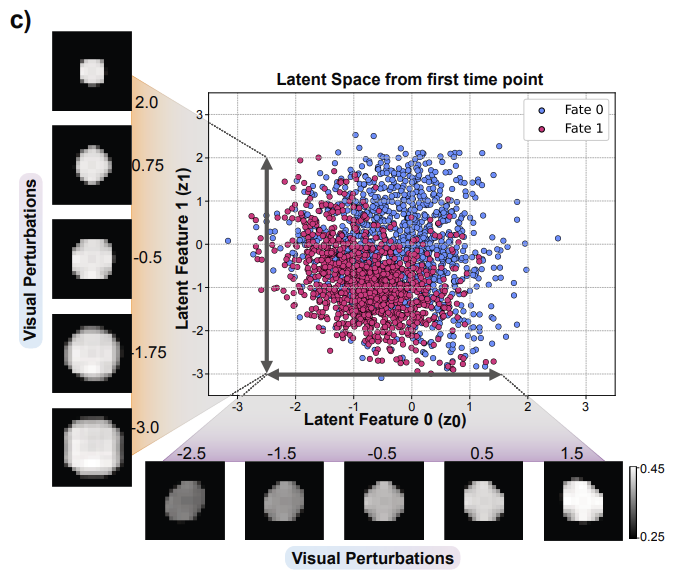
While Gilead focused on accelerating the existing workflow, the AI4CellFate [1] framework, introduced by researchers at the Science for Life Laboratory, suggests we can look even deeper. Using generative AI and contrastive learning, this model aims to predict a cell’s “fate”—whether it will proliferate or fail—from the very first frame of a microscopy timelapse.
AI4CellFate [1]: This digital cartography reduces a cell’s messy reality into a two-dimensional “latent space,” where its biological destiny is written in its physical traits. By applying visual perturbations, the AI4CellFate model demystifies its own logic: the horizontal axis (z_0) tracks signal intensity—a proxy for ERK activity—while the vertical axis (z_1) measures cell size. Proliferating cells (Fate 1, in red) gravitate toward a specific neighborhood of higher intensity and larger size, allowing the machine to predict future growth from the very first frame of imaging. This move beyond the “black box” provides an interpretable, data-driven platform for identifying the industry’s most valuable clones.
The implications are transformative. The research suggests that the signatures of future success—stable signaling and specific morphological traits—are present long before a cell actually divides. By engineering a “latent space” that mathematically disentangles features like cell size and signal intensity, AI4CellFate provides an unbiased platform for understanding why a cell is destined for greatness. This moves the industry beyond hypothesis-driven markers toward a purely data-driven understanding of cell health.
The Morphological Convergence
This trend is further supported by recent industry findings, such as those from Sartorius [3], which demonstrate that high-producing clones can be identified within just four days of cloning simply by analyzing morphology. Clones selected via these AI-driven morphological insights showed up to 60% higher titers than those chosen through traditional manual ranking.
The common thread is clear: cell morphology is no longer just a visual check; it is a digital biomarker for productivity and fate.
Closing the Loop
As these AI models become more sophisticated, the challenge shifts from observing these traits to acting on them in real-time. This is the catalyst for Deepcell’s recent collaborations with various players across the biopharma space.
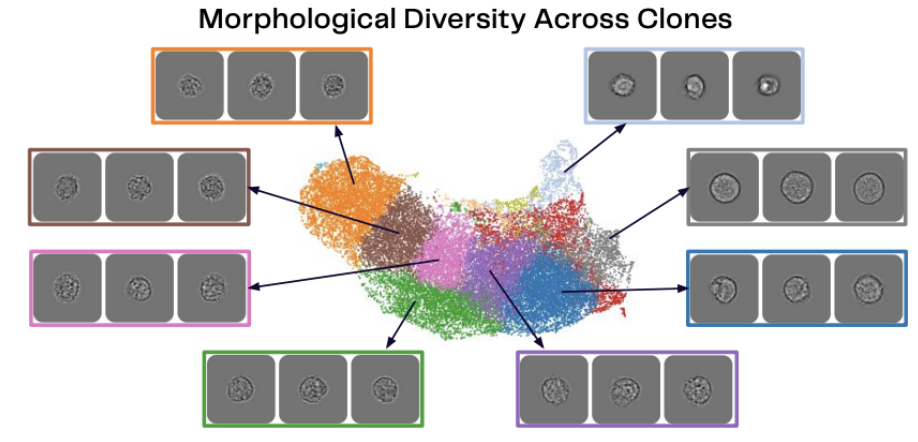
This high-dimensional map [4] made using Deepcell REM-I reveals the hidden heterogeneity within a single CHO cell clone, a phenomenon that has long complicated biologics manufacturing.
While tools like CLAIRE and AI4CellFate provide the “vision” to recognize high-potential clones, the industry requires a platform that can combine this high-resolution imaging and AI analysis with the physical ability to
sort and capture these cells as they flow through a system. By positioning high-dimensional AI at the point of sorting, the goal is to turn “Day 0” predictions into immediate, actionable results.
In 2026, the question is no longer whether we can see a cell’s future, but how quickly we can capture it.
References
AI4CellFate: Interpretable Early Cell Fate Prediction with Generative AI. Cunha, I., Panconi, L., et al. bioRxiv (2025). https://doi.org/10.1101/2025.05.12.653464
An AI/ML-Powered Workflow for End-to-End Cell Line Development. Unnikandam Veettil, S., Donatelli, J., et al. bioRxiv (2026). https://doi.org/10.64898/2026.02.04.703387
How AI Is Transforming Cell Line Development for Protein Production. Sartorius Cell Line Development Center of Excellence. Science Snippets (2026). https://www.sartorius.com/en/knowledge/science-snippets/ai-in-cell-line-development-1757086
Morphological Diversity Across Clones. Deepcell Internal Data / Technical Showcase (2026).